 
"I believe that the integration of scientific tasks into established computer games will be a commonly used approach in the future to harness the brain processing power of humans, and that intricate designs of citizen science games feeding directly into machine learning models has the power to rapidly leverage the output of large-scale science efforts," Lundberg says. By combining players'annotations with the machine learning approach, transfer-learning was used to boost the performance of Loc-CAT significantly. 
Loc-CAT is the first generalized tool for annotating proteins with multiple localizations from images and generalizes across a large number of cell types, providing a useful tool in studying cells and their behavior in the future.Īlthough Loc-CAT outperformed players of Project Discovery for many of the common classes of proteins, aggregated player data from EVE Online better identified rare classes and was able to annotate new patterns where training data was unavailable. The ability of citizen scientists was compared to an AI system for predicting protein subcellular localization from images called the Localization Cellular Annotation Tool (Loc-CAT). It was played by more than 300,000 citizen scientists within EVE Online who together generated more than 33 million image classifications of protein subcellular localization, an achievement hailed as a milestone in citizen science. The resulting mini-game called "Project Discovery" featured Lundberg's avatar, making her one of the first living scientists to be featured in an online game. 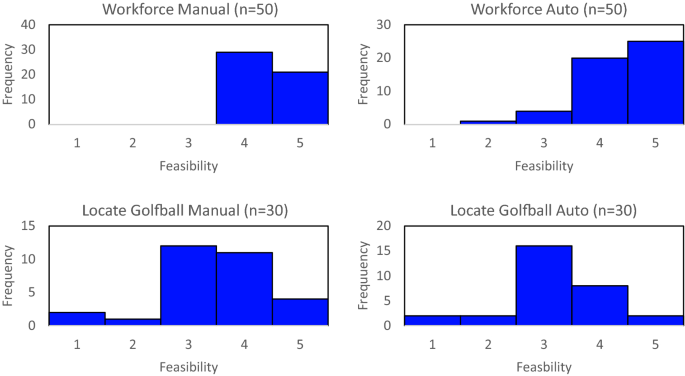
The researchers partnered with Massively Multiplayer Online Science and CCP Games to integrate analysis of protein localization from the Human Protein Atlas Cell Atlas images directly into EVE Online, a popular massively multiplayer online game. Lundberg says the data is being actively integrated into the publicly-available Human Protein Atlas database and will be a resource for researchers worldwide who are working towards a greater understanding of human cells, proteins and disease development. The combination of crowdsourcing and AI led to improved classification of subcellular protein patterns and the first-time identification of 10 new members of the family of cellular structures known as "Rods & Rings," says Emma Lundberg, a researcher from KTH who leads the Cell Atlas, part of the Human Protein Atlas, at the Science for Life Laboratory joint research center. In a study published in the September issue of Nature Biotechnology, the researchers found that gamers, or "citizen scientists," helped boost the AI system used for predicting protein localization on a subcellular level. The advances were reported by a collaboration between KTH Royal Institute of Technology, CCP Games and Massively Multiplayer Online Science. Results from a groundbreaking citizen science projectīuilding on a map that shows hundreds of thousands of microscopic images of human cells, an international research team is working with the gaming community and with artificial intelligence to gain a more granular understanding of patterns of proteins arranged within the body's cells. Mapping of cells and proteins improved with combination of massively multiplayer crowdsourcing and AI
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Blog
- Home
- Free Urdu Guide To Dairy Farm
- Download Forza 3 Pc Full Version Free
- Hopper 3
- Jodha Akbar Hindi Audio Songs Free Download
- Jude Full Hindi Dqonlode
- Basara 4 Pc Crack
- Bartender 3.0.12 Crack Plus License Key For Mac
- File Synchronization Via Soap Over Http
- Nokia Universal Imei Changer 1.0 Rar Gsm
- U167controller Usb Device Driver For Mac
- Download Video Alif Ba Ta Song
- Underground Vibes Rar Extractor
- Html5 Plugin For Firefox
- Bharathidasan Poems In Tamil Pdf Hot
- Kumpulan Template Zw Terbaru
- Arabic Song Mp3 UAE
- 3d Avatar Creator No
- The Datsuns Rar
- Durga Tarang Tv Serial All Artist Real Name
- Pes 6 Serial Crack
- How To Cook Up Crack With Ammonia
- Boris Vian Les Fourmis Pdf Editor
- Snapper Serial Number Years
- Installing Porclean Tile
 RSS Feed
RSS Feed
